|
ABE is a Java-based standalone program to annotate and analyze proteomic and transcriptomic data sets.
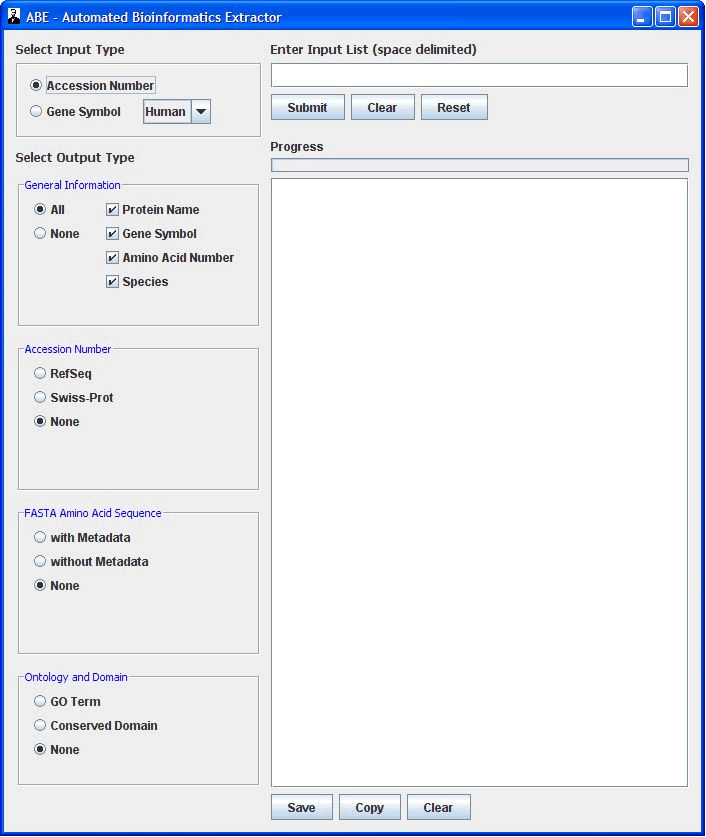
ABE uses protein accession number or gene symbol to extract bioinformatic data outlined in the below table.
| Input |
Output |
accession number
or
gene symbol
|
General information: protein name, gene symbol, amino acid number, and species
|
| Accession number: RefSeq or Swiss-Prot |
| FASTA sequence with or without metadata |
| GO term |
| Conserved domain |
Download ABE here: ABE.jar
How to start ABE:
- Make sure that you have the most recent Java Platform. If not, download it from this link http://www.java.com/en/download/index.jsp
- Prepare a single column input list in Excel (eliminate blank space, if any). Input can be either accession number or gene symbol. Different types of accession number can be used, however, some accession number types can not be used to extract GO term and conserved domain, see the below table.
| Accession Number |
Output |
General
information |
Accession
number |
FASTA
sequence |
GO term |
Conserved
domain |
| RefSeq (ref) |
+ |
+ |
+ |
+ |
+ |
| Swiss-Prot (sp) |
+ |
+ |
+ |
- |
- |
| GenBank (gb) |
+ |
+ |
+ |
+ |
+ |
| EMBL (emb) |
+ |
+ |
+ |
+ |
+ |
| GenInfo Identifier (gi) |
+ |
+ |
+ |
- |
- |
- Copy and paste the input list into input text field.
- Select input: accession number or gene symbol. Select species (human, rat, or mouse) if gene symbol is used.
- Select output. More than one type of output can be selected among general information, accession number, and FASTA sequence. Selection of either GO term or conserved domain will inactivate other output types, to resume select “None” button in Ontology and Domain panel.
- Click "Submit" button to start extracting data.
- When the extraction is done you can either copy data onto clipboard using “Copy” button or save data into a file using "Save" button.
ABE was created by Dmitry Tchapyjnikov, Patricia Gonzales, Jason D. Hoffert, Mark A. Knepper, and Trairak Pisitkun.
Contact us with questions or comments.
|