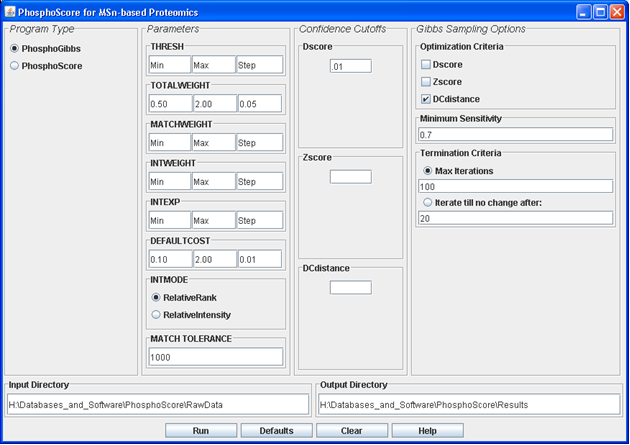
PhosphoScore is a phosphorylation assignment program that is compatible with all levels of tandem mass spectrometry spectra (MSn) generated through the Bioworks/Sequest platform. The program utilizes a “cost function” which takes into account both the match quality and normalized intensity of observed spectral peaks compared to a theoretical spectrum.
Technical Details -
PhosphoScore was written in Java. The program is accessible through a graphical user interface (GUI) or can be run from the command line. The input directory should contain all Sequest DTA and OUT files as well as the Sequest parameters (sequest.params) file from the original LC-MSn analysis. Data from different MS levels should be analyzed separately by PhosphoScore due to differences in Sequest search parameters. Please consult the "readme.txt" file for more details on running the program. For a complete description of the software, please see
Ruttenberg et al. J Proteome Res. 2008 Jul;7(7):3054-9.
Files for download -
code.zip Includes the PhosphoScore program, the Java source code, and the "readme.txt" file that will help get you started.
sampledata.zip A small set of MS2 and MS3 dta/out files along with their corresponding "sequest.params" files that you can test before you try your own data.
PhosphoScore was created by Brian E. Ruttenberg, Trairak Pisitkun, Mark A. Knepper, and Jason D. Hoffert.
Contact us with questions.
|